The intestinal microbiota is significantly altered in cirrhotic patients, but the composition of the intestinal microbiota in Mexican patients with the pathology has not been reported. The present study is an attempt to determine the type of intestinal microbiota in healthy subjects and in patients of Mexican mestizo origin that present with cirrhosis of the liver.
Materials and methodsBiochemical liver function parameters (ALT, AST, GGT, BIL-T, etc.) were determined in 23 cirrhotic patients and 21 control subjects. The intestinal microbiota was established through 16S ribosomal RNA gene sequencing.
ResultsThe cirrhotic patients had elevated levels of ALT, AST, GGT (105.2±77.7 vs. 20.99±8.5UI/L, 110±68.6 vs. 23.39±5.2, and 119.1±79.1 vs. 19.3±15.2UI/L, respectively), IL-6 (1.64±0.38pg/ml, P<.001), or TNFα (1.78±0.3, P<.05). The intestinal microbiota of the cirrhotic patients was less diverse, compared with that of the control subjects. At the level of the phylum, there was a significant increase in Proteobacteria and Bacteroidetes in the patients with cirrhosis, compared with the controls (6.2 vs. 4.9% and 44 vs. 46%, respectively, P<.01). In contrast, there was a decrease in Firmicutes, Actinobacteria, and Fusobacteria in the cirrhotic patients. There was an increase in the Campylobacter and Gemella families in the cirrhotic patients, whereas Streptococcus and Veillonella had a positive association with serum ALT or AST levels.
ConclusionsTo the best of our knowledge, the present study is the first to demonstrate the type of intestinal microbiota in Mexican patients with cirrhosis of the liver. The extension of the findings in a larger cohort of subjects and the metagenome analysis will enable the creation of data that can have relevant treatment implications for this group of patients in Mexico.
La microbiota intestinal se altera significativamente en pacientes cirróticos. Sin embargo, la composición de la microbiota intestinal en pacientes cirróticos mexicanos no ha sido reportada. En este trabajo nos propusimos determinar el tipo de microbiota intestinal en sujetos sanos y pacientes con cirrosis hepática de origen mestizo-mexicano.
Material y métodosSe determinaron los parámetros bioquímicos de la función hepática (ALT, AST, GGT, BIL-T, etc.) en 23 pacientes cirróticos y 21 sujetos control. Se determinó la microbiota intestinal mediante secuenciación del gen ribosomal 16S.
ResultadosLos pacientes cirróticos presentan niveles elevados de ALT, AST, GGT (105.2±77.7 vs. 20.99±8.5UI/L; 110±68.6 vs. 23.39±5.2 y 119.1±79.1 vs. 19.3±15.2UI/L, respectivamente) e IL-6 (1.64±0.38pg/ml, p<0.001) o TNFα (1.78±0.3, p<0.05). La microbiota intestinal de los pacientes cirróticos es menos diversa comparada con sujetos control. A nivel de phylum, las Proteobacteria y los Bacteroidetes aumentan significativamente en cirróticos (6.2 vs. 4.9% y 44 vs. 46% vs. controles, respectivamente, p<0.01). En contraste, los Firmicutes así como Actinobacteria y Fusobacteria, disminuyeron en los pacientes cirróticos. Las familias Campylobacter y Gemella aumentan en los pacientes cirróticos, mientras que Streptococcus y Veillonella se asocian positivamente con los niveles séricos de ALT o AST.
ConclusionesDemostramos por vez primera el tipo de microbiota intestinal en pacientes cirróticos mexicanos. La extensión de los hallazgos en una cohorte más amplia de sujetos y el análisis del metagenoma permitirán generar datos que pueden tener implicaciones relevantes para el tratamiento de este grupo de pacientes en México.
Cirrhosis of the liver is characterized by the presence of chronic inflammatory processes, portal hypertension, and alterations in the metabolism of fats, carbohydrates, and ammonium, among others. In addition, it can cause cognitive damage in the long term.1 It has been reported that liver damage from cirrhosis can reduce the flow of blood that travels through the gut-liver axis, altering bile acid secretion and intestinal peristalsis, which in turn promotes mucosal barrier damage and possibly gut microbiota alteration.2 Under normal conditions, the gut microbiota mainly consists of the phyla Firmicutes (60%) and Bacteroidetes (15-20%), whereas Actinobacteria (< 10%) and Proteobacteria (1%) are less abundant.3 However, that composition can change in response to different illnesses, such as liver disease. Previous studies indicate that cirrhotic patients present with increases in the phyla Fusobacteria and Proteobacteria, as well as the Enterobacteriaceae, Veillonellaceae, and Streptococcaceae families, whereas there is a decrease in Bacteroidetes and Actinobacteria with respect to healthy patients.4–6 In a North American population, patients with cirrhosis due to alcohol consumption or steatohepatitis (nonalcoholic steatohepatitis) were observed to present with changes in the composition of the gut microbiota, depending on disease stage. In patients with compensated or decompensated cirrhosis an increase in the number of Bacteroidetes has been seen, compared with healthy patients,1 whereas hospitalized patients have shown a reduction of that phylum.7 Additional evidence indicates that high levels of the families Enterobacteriaceae, Alcaligenaceae, and Fusobacteriaceae are found in cirrhotic patients, whereas levels of Ruminococcaceae and Lachnospiraceae are reduced, compared with control subjects.8 Thalheimer et al. posited the idea that the microbiota, or at least some of the bacteria it is composed of, can compensate for the loss of metabolic capacity in patients affected by cirrhosis of the liver derived from hepatitis B virus infection. They described a negative or positive correlation between the phylum Bacteroidetes or the families Enterobacteriaceae and Veillonella, respectively, and the Child-Pugh score, and the use of those markers as factors for distinguishing the microbiota of patients with cirrhosis from that of healthy subjects.9
On the other hand, bacterial translocation is an important risk factor for the development of certain complications of cirrhosis of the liver, such as spontaneous bacterial peritonitis, hepatic encephalopathy, and hepatorenal syndrome.9,10 Bacterial translocation in patients with cirrhosis has been corroborated through bacterial DNA quantification in blood and ascitic fluid.11,12
The metabolic and physiologic effects resulting from those changes in the gut microbiota composition in patients with cirrhosis of the liver is an area of interest. However, the majority of studies that characterize the gut microbiota in cirrhotic patients have been conducted on populations in Asia and the United States. No data are available yet on the characterization of the microbiota in healthy persons and cirrhotic patients in the Mexican population. The determination of those characteristics will be of great usefulness for establishing the differences and similarities between the Mexican mestizo populations and the other populations studied. Thus, the primary aim of our study was to determine the patterns of the gut microbiota in a Mexican mestizo population of healthy subjects and patients with cirrhosis of the liver, as well as to identify the changes in the abundance of the bacteria that make up the gut microbiota of patients with cirrhosis of the liver and correlate them with the parameter of liver damage.
Materials and methodsThe present study was approved by the Ethics and Research Committee of the Instituto Nacional de Ciencias Médicas y Nutrición Salvador Zubirán. All patients were given information as stipulated in the guidelines established by that committee and they provided their written statements of informed consent.
A cross-sectional case-control study was conducted. Twenty-one healthy subjects and 23 patients with compensated cirrhosis of the liver, whose clinical case records were reviewed by an expert hepatologist, were recruited for the study. Only patients with a Child-Pugh score ≤ 5 points, equivalent to class A,13 were included. Patients with unconfirmed diagnosis of cirrhosis and healthy subjects whose ultrasound study of the liver revealed grade 1 fatty liver, or higher, were excluded from the study, as well as patients taking probiotics, prebiotics, antibiotics, antifungal agents, antivirals, or steroids. Also excluded were patients consuming any abusive drug or alcohol for more than 8 weeks before the study that could alter microbiota characteristics, in addition to subjects with morbid obesity, type 2 diabetes, chronic bowel disease, cancer, or HIV or hepatitis A infection. Anthropometry was evaluated through bioelectric impedance with 8-point tetrapolar tactile electrodes to determine body composition utilizing the InBody 770 (InBody, Korea) device. Stool samples were both collected and refrigerated the morning prior to delivery.
Blood metabolite determinationComplete liver function tests (ALT, AST, GGT, albumin, creatinine, total bilirubin, direct bilirubin, indirect bilirubin, triglycerides, and total cholesterol) were determined through the automated Cobas C111 (Roche, USA) equipment. Serum proinflammatory cytokine TNFα, IL-6, IL-8, IL-10, and IL-12 expression was determined using the magnetic ELISA kit (LuminexAssay, R&D, USA), and LPS levels were determined through the Limulus amebocyte lysate (QCL-1000™ Assay, Lonza USA) test.
Stool genomic DNA extractionStool genomic DNA extraction was performed according to the instructions of the commercial QIAamp Fast Stool DNA Mini Kit (Qiagen, USA). Genomic DNA was stored at −70°C until its processing for the next-generation sequencing stage.
16S rRNA gene sequencing and data analysisThe genomic DNA obtained was quantified utilizing the Qubit® dsDNA HS Assay reagent, (Molecular Probes, USA). An end-point amplicon was produced through PCR and the oligonucleotides reported for the hypervariable V3-V4 region of the 16S rRNA bacterial gene were obtained.14 The amplicons were used to produce the individual index libraries that later were normalized and grouped for loading in the NextSeq (Illumina, USA) flow cell. The data were analyzed using the MICCA (Microbial Community Analysis, GPL v3) program15 and the QIIME v 1.9.1. program.16 The LEfSe program in the Galaxy Work Framework platform17 was utilized for establishing the taxa associated with the healthy subjects or the cirrhotic patients.
Statistical analysisThe analysis of anthropometric data, biochemical tests, and cytokine and LPS testing were carried out using the Prism7 (Graph-Pad, EUA) program, with the unpaired Student's t test. Statistical significance was set at a p < 0.05. Conglomerates for comparing the anthropometric and biochemical data, interleukins or LPS, and the relative bacterial abundance (operational taxonomic unit [OTU]), were created with the data analyzed in the SPSS program (IBM, v21, USA), utilizing Pearson correlations. The difference in bacterial abundance (alpha-diversity) between the two groups was compared using the two-tailed Student's t test, with the Shannon and Chao1 indexes. The analyses of the correlation between the biochemical variables and the OTUs were carried out using the SPSS (IBM v21, USA) program, whereas the correlations between the different taxa were obtained using the MEGAN v618 and SPSS v 21 (IBM, USA) programs.
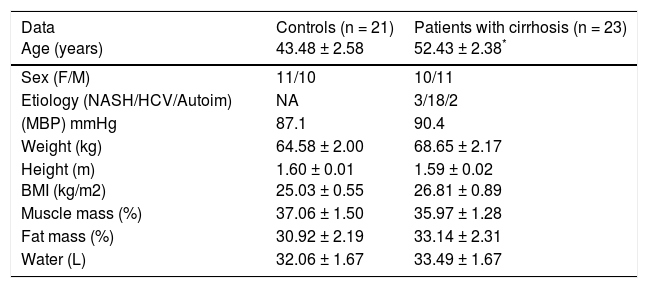
ResultsAnthropometric and biochemical characteristics of the participantsA total of 44 patients were recruited: 23 patients with cirrhosis and 21 healthy controls. Of all the cirrhotic patients, the causes were nonalcoholic steatohepatitis in 3 (13%) of them, autoimmune disease in 2 (8.7%), and C virus infection in 18 (78%). The clinical and anthropometric characteristics of the two groups are presented in Tables 1 and 2. The data indicated that the liver damage enzymes of ALT, AST, GGT, and AP were significantly high in the patients with cirrhosis, compared with the control subjects (Table 2).
General characteristics of the participants.
| Data Age (years) | Controls (n = 21) 43.48 ± 2.58 | Patients with cirrhosis (n = 23) 52.43 ± 2.38* |
|---|---|---|
| Sex (F/M) | 11/10 | 10/11 |
| Etiology (NASH/HCV/Autoim) | NA | 3/18/2 |
| (MBP) mmHg | 87.1 | 90.4 |
| Weight (kg) | 64.58 ± 2.00 | 68.65 ± 2.17 |
| Height (m) BMI (kg/m2) | 1.60 ± 0.01 25.03 ± 0.55 | 1.59 ± 0.02 26.81 ± 0.89 |
| Muscle mass (%) | 37.06 ± 1.50 | 35.97 ± 1.28 |
| Fat mass (%) | 30.92 ± 2.19 | 33.14 ± 2.31 |
| Water (L) | 32.06 ± 1.67 | 33.49 ± 1.67 |
Autoim: autoimmune; BMI: body mass index; HCV: hepatitis C virus; MBP: mean blood pressure; NA: not applicable; NASH: non-alcoholic steatohepatitis.
Data are presented as mean ± SD.
Biochemical parameters of the participants.
| Parameter | Controls (n=21) | Patients with cirrhosis (n=23) | p |
|---|---|---|---|
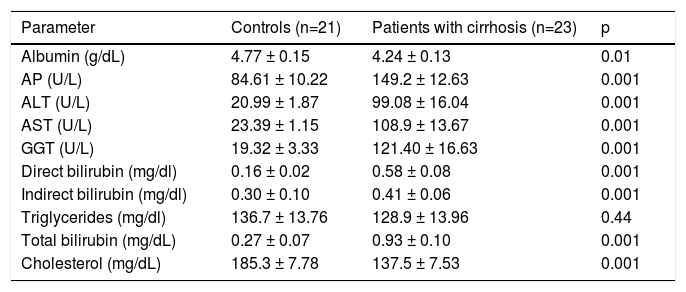
| Albumin (g/dL) | 4.77 ± 0.15 | 4.24 ± 0.13 | 0.01 |
| AP (U/L) | 84.61 ± 10.22 | 149.2 ± 12.63 | 0.001 |
| ALT (U/L) | 20.99 ± 1.87 | 99.08 ± 16.04 | 0.001 |
| AST (U/L) | 23.39 ± 1.15 | 108.9 ± 13.67 | 0.001 |
| GGT (U/L) | 19.32 ± 3.33 | 121.40 ± 16.63 | 0.001 |
| Direct bilirubin (mg/dl) | 0.16 ± 0.02 | 0.58 ± 0.08 | 0.001 |
| Indirect bilirubin (mg/dl) | 0.30 ± 0.10 | 0.41 ± 0.06 | 0.001 |
| Triglycerides (mg/dl) | 136.7 ± 13.76 | 128.9 ± 13.96 | 0.44 |
| Total bilirubin (mg/dL) | 0.27 ± 0.07 | 0.93 ± 0.10 | 0.001 |
| Cholesterol (mg/dL) | 185.3 ± 7.78 | 137.5 ± 7.53 | 0.001 |
AP: Alkaline phosphatase; ALB: Albumin; ALT: Alanine-aminotransferase; AST: Aspartate-aminotransferase; GGT: gamma glutamyl-transferase.
Data are presented as mean ± SD.
*p < 0.05; **p < 0.01; ***p < 0.001
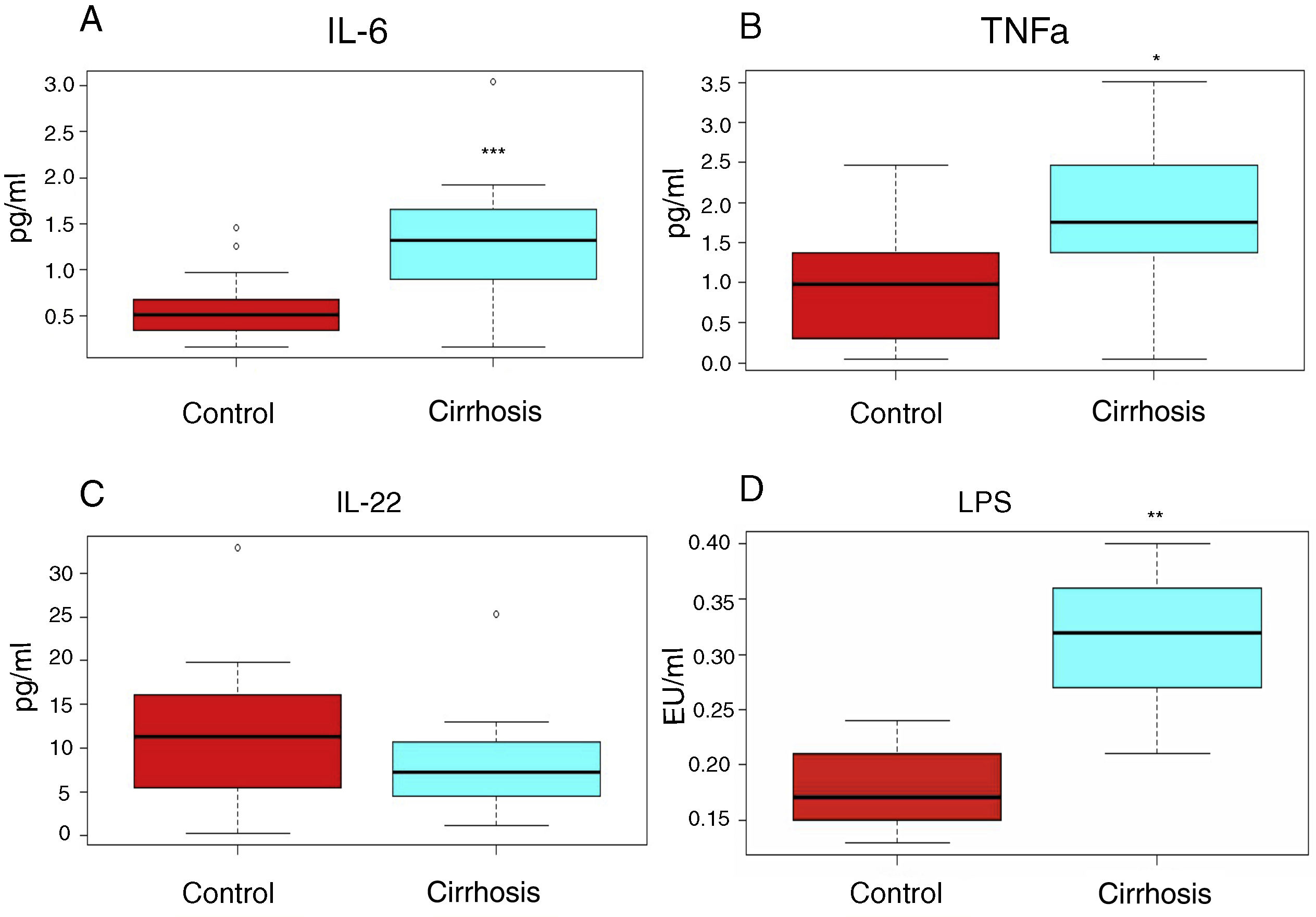
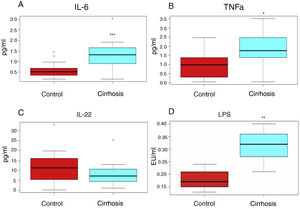
In addition, the proinflammatory cytokine levels (IL-6, TNF-alpha) were high (1.64 ± 0.38 pg/ml vs. 0.558 ± 0.08 pg/ml, 1.78 ± 0.29 pg/ml vs. 0.94 ± 0.21pg/ml, respectively), whereas the levels of the anti-inflammatory interleukin, IL-22, were below those observed in the control subjects (8.32 ± 1.53 pg/ml vs. 11.96 ± 2.55pg/ml) (fig. 1A-C).
The cirrhotic patients presented with a higher inflammation grade, compared with the control subjects. The interleukins (figs. 1 A, 1 B, 1C) or lipopolysaccharides (LPS) (fig. 1 D), determined in the peripheral blood serum of the cirrhotic patients (blue boxes) or control subjects (red boxes) are shown. The graphs show the medians and interquartile ranges. The asterisks indicate statistical significance ***p < 0.001; **p < 0.01; *p < 0.05 vs. control.
LPS levels can be used as mild-to-moderate endotoxemia indicators, and in turn, as the grade of inflammation markers in patients with cirrhosis, and they were elevated in our cirrhotic patients, compared with those of the healthy subjects (fig. 1D).
Together, those data indicated that our population of patients with cirrhosis presented with inflammation that could be related to bacterial translocation.
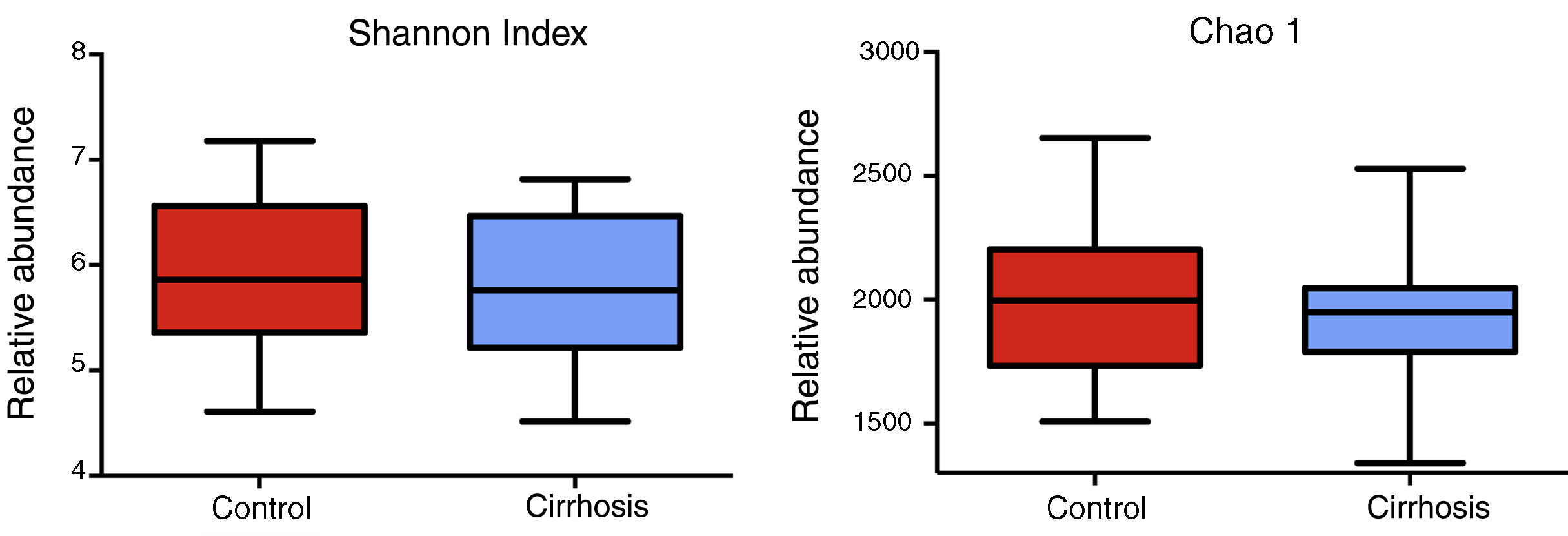
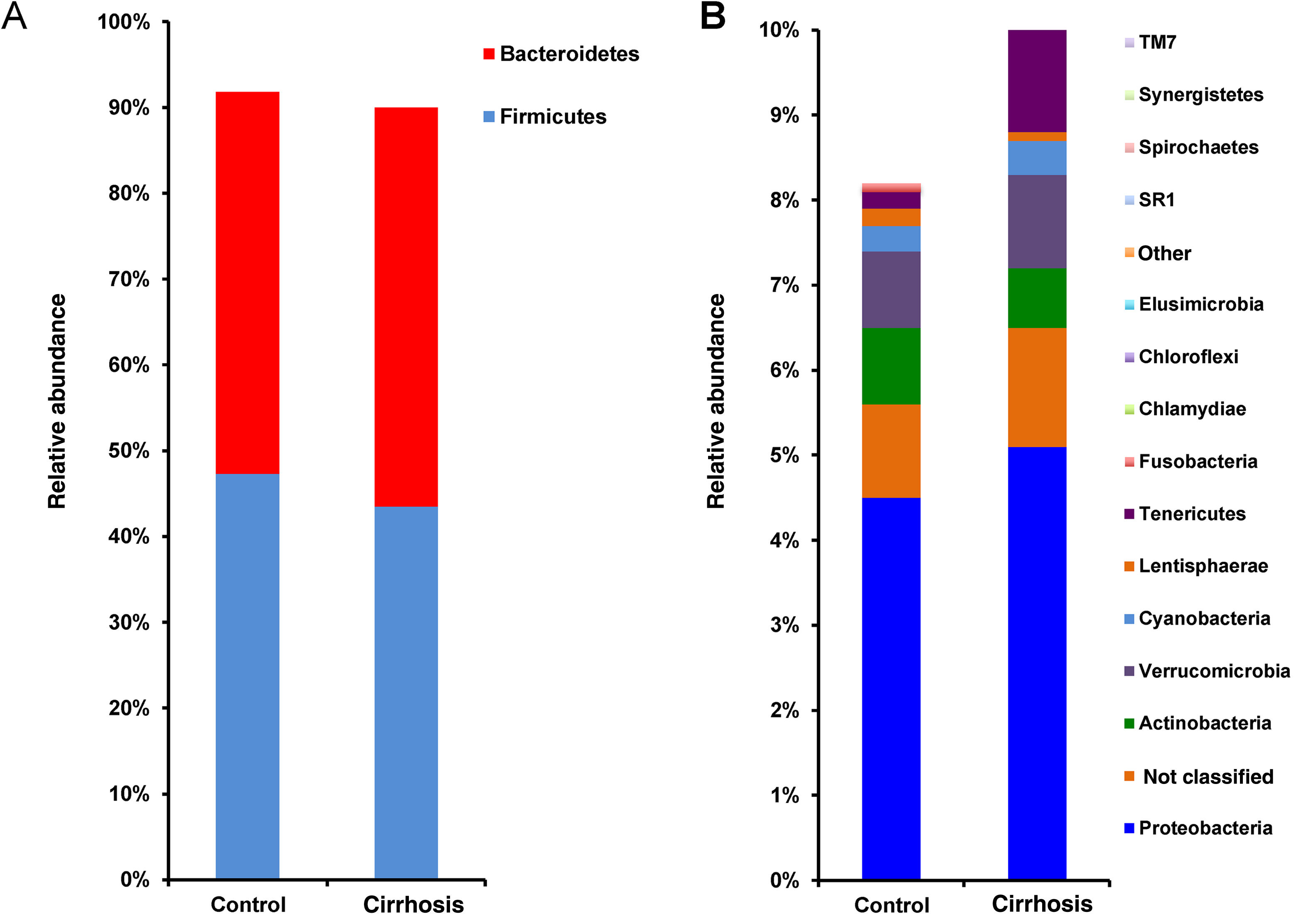
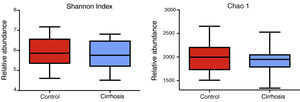
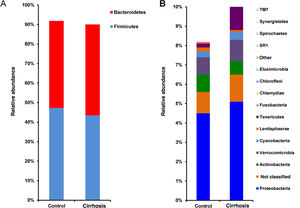
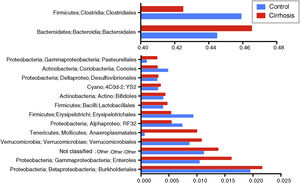
Composition and functional analysis of the microbiota of the patients with cirrhosisThe diversity analyses performed through the Chao1 and Shannon indexes, comparing the cirrhotic subjects and the controls, showed less bacterial diversity in the patients with cirrhosis. Even though that difference did not reach statistical significance, it did indicate that the microbiota of the cirrhotic patients was directly or indirectly affected by the disease (fig. 2). The analysis of the taxonomic frequencies in each group revealed that the patients with cirrhosis had a higher number of OTUs belonging to the phyla Bacteroidetes, Proteobacteria, Verrucomicrobia, Cyanobacteria, and Tenericutes, as well as a lower abundance of OTUs in the phyla Actinobacteria, Fusobacteria, Lentisphaerae, Synergistetes, TM7, SR1, and Spirochaetes, among others (fig. 3A, 3B).
The cirrhotic patients showed less bacterial diversity, compared with the control subjects. The alpha-diversity of the two groups (44 patients) is demonstrated through the Shannon and Chao1 indexes. The graphs describe the interquartile ranges between the first and third quartiles (25th and 75th percentiles) and the line dividing the box represents the median of each group. Shannon p < 0.58; Chao1 p < 0.29 vs. control, Mann-Whitney test.
Comparison of the relative abundance of the OTUs at the phylum level between the cirrhotic patients and the control subjects (fig. 3A). The superimposed bars of the graph indicate the mean of the most abundant phyla and present higher levels of change ± standard error, in each case (fig. 3B). As in A, the bars indicate the levels of change.
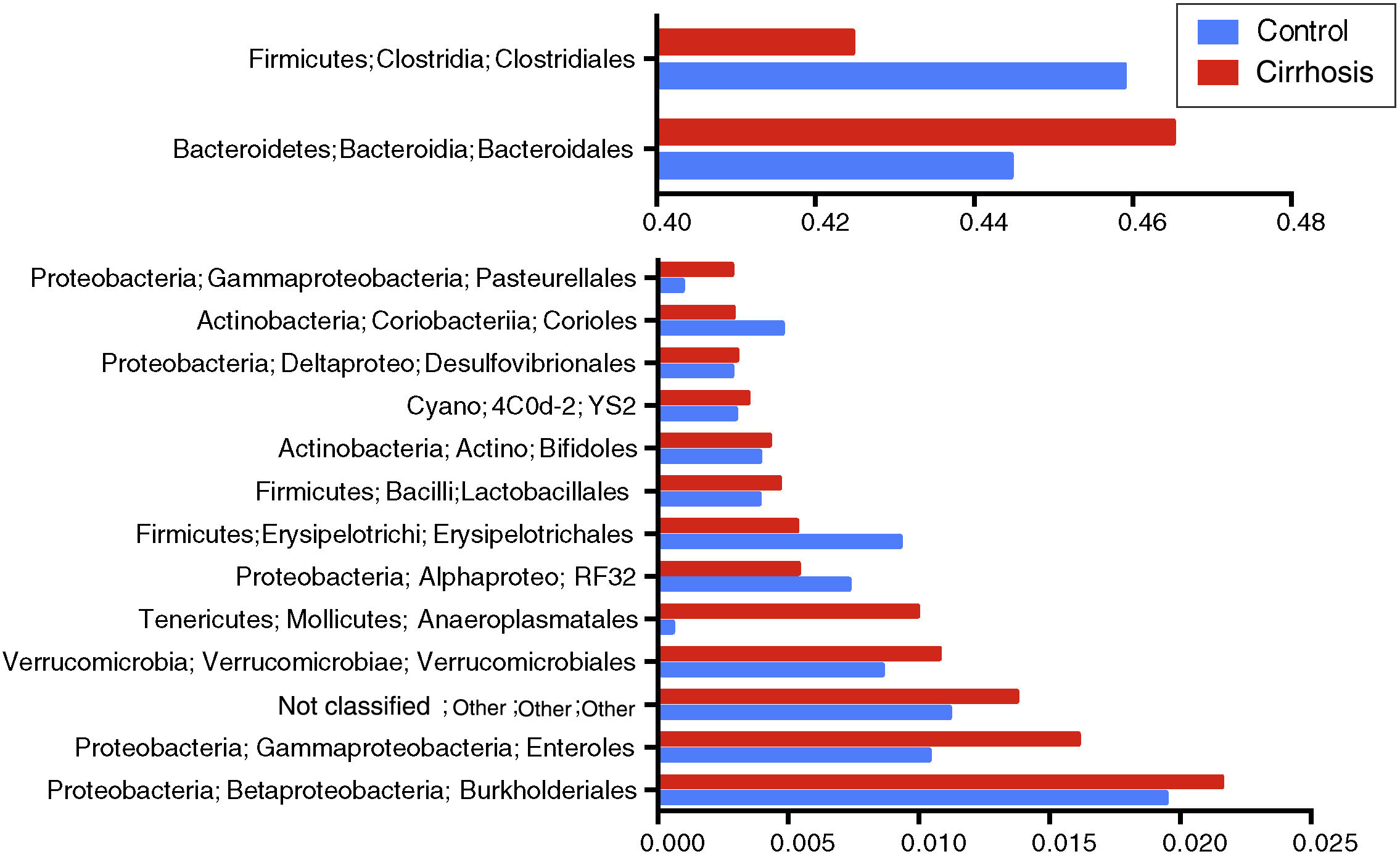
At the level of order, we found that Bacteroidales and Anaeroplasmatales, Verrucomicrobiales, and Enterobacteriales, as well as the Burkholderiales and a group of unclassified bacteria dominated the gut microbiota of the cirrhotic patients, whereas Clostridiales, Erysipelotrichales, and Coriobacteriales were dominant in the microbiota of the control subjects (fig. 4).
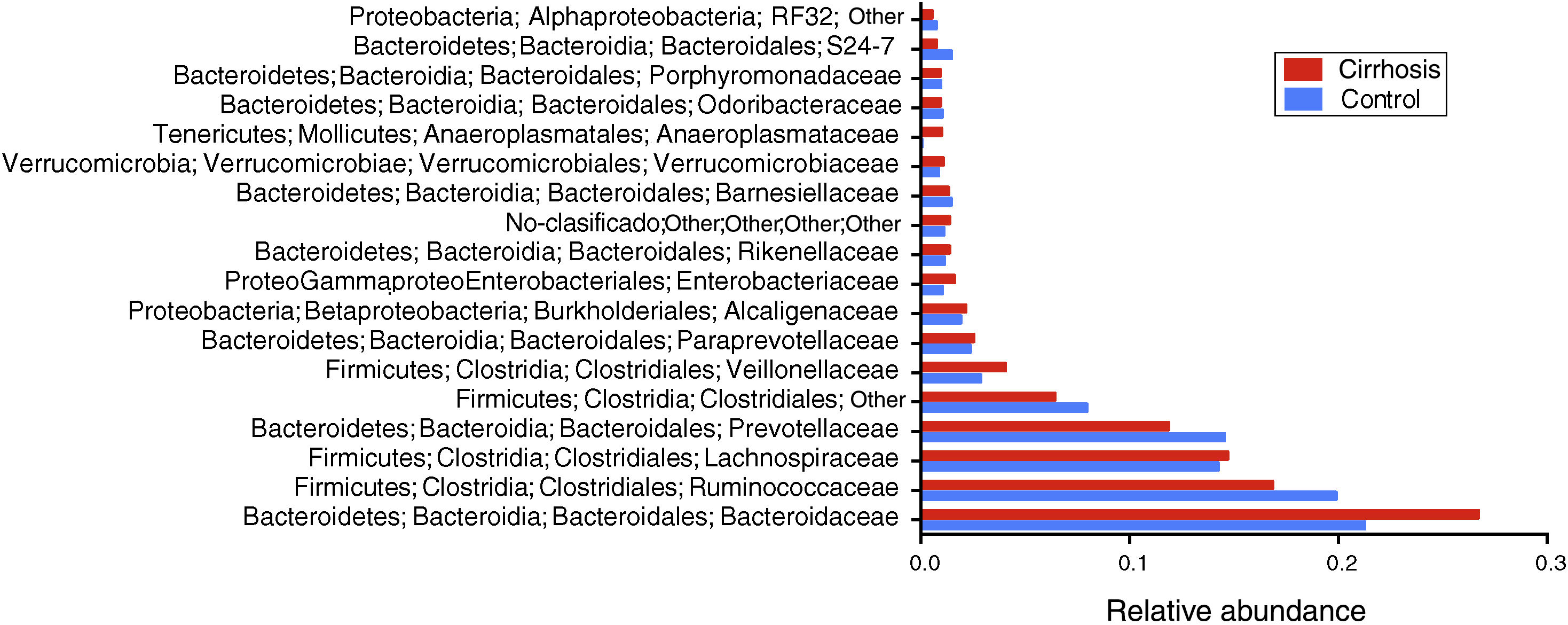
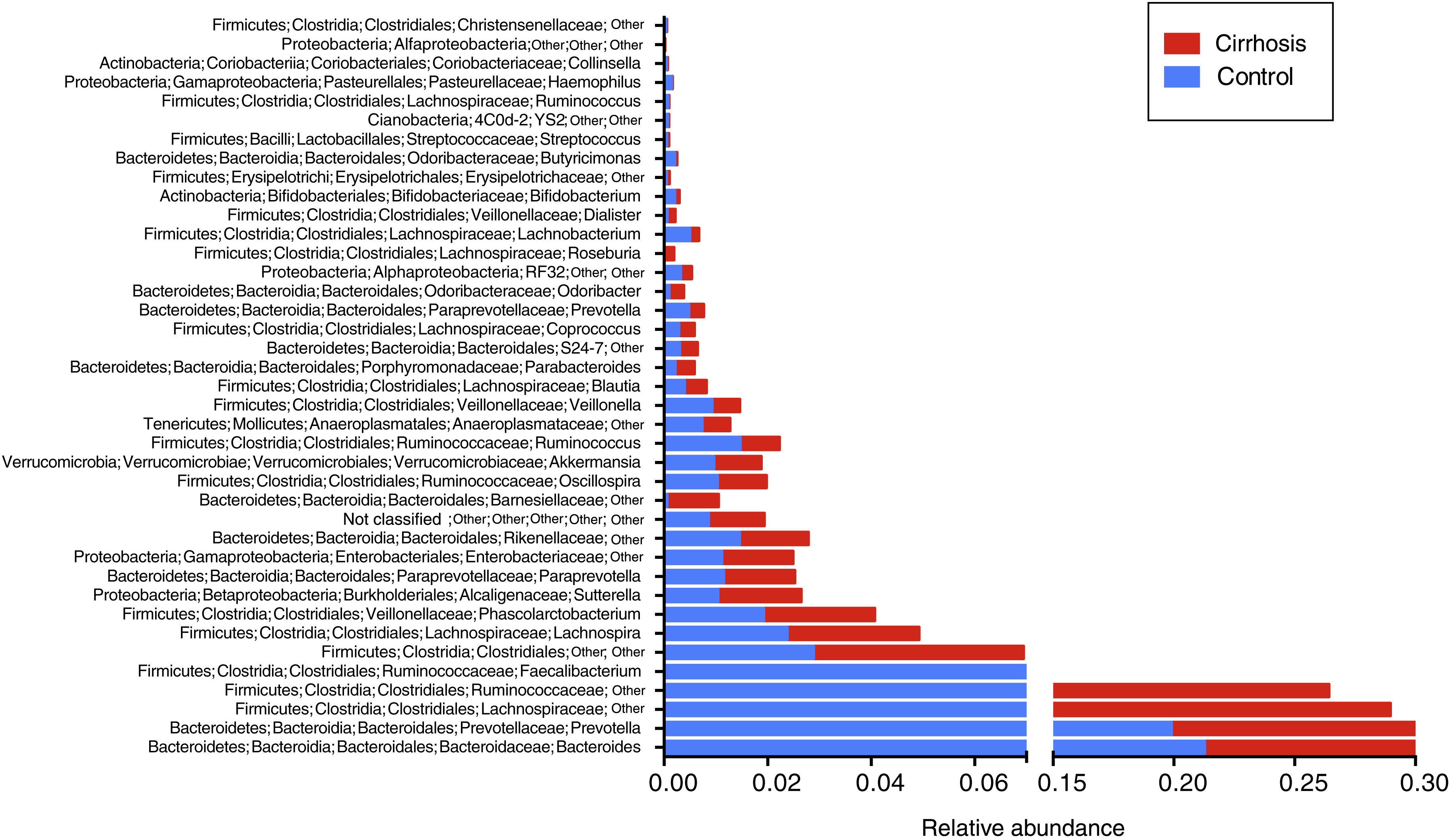
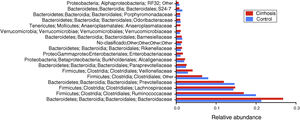
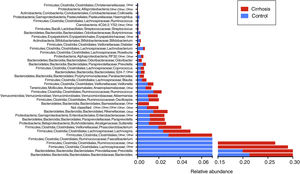
When the gut microbiota analysis was carried out at the level of family and genus, we found that the families Bacteroidaceae, Enterobacteriaceae, Rickenellaceae, and Veillonellaceae were augmented in the patients with cirrhosis, compared with the control group (fig. 5), whereas the genera that were elevated in the cirrhotic patients were Bacteroides, Phascolarctobacterium, Sutterella, Paraprevotella, Veillonella, and Blautia. On the other hand, we found that Prevotella, unclassified Ruminococcaceae, unclassified Clostridiales, as well as Lachnospiraceae, were more abundant in the control subjects (fig. 6).
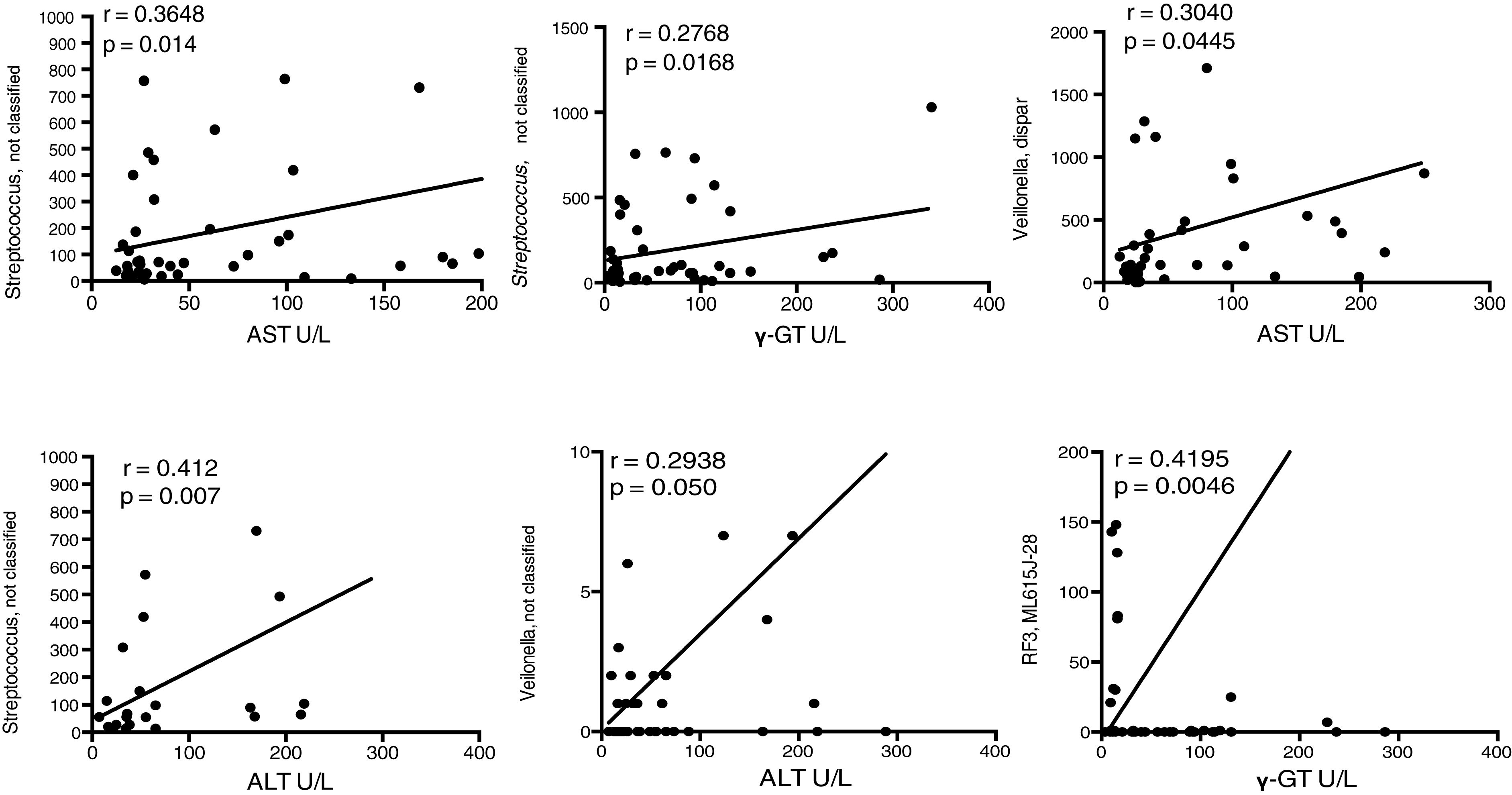
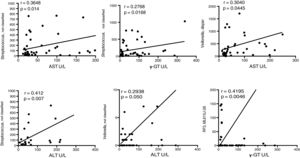
For the purpose of establishing associations between families of bacteria in the patients with cirrhosis and the liver damage indicators, we decided to carry out the correlations of the biochemical alterations with the changes in OTU abundance in the two groups. In that analysis, several OTUs had positive correlations with the cirrhotic patients (fig. 7), indicating that the changes in those bacterial groups, specifically the increase we found in their relative abundance, could be related to the initiation or progression, or perhaps both, of liver damage, mainly due to C virus infection.
Fragment of the bacterial signatures that present significant associations with liver damage etiology. The graphs indicate the levels of association between OTUs at the family level and serum levels of the enzymes that indicate liver damage. The correlation curve and the r-squared, as well as the significance of the Pearson test are presented in each case. ALT: alanine aminotransferase; AST, aspartate aminotransferase; γ-GT, gamma-glutamyl transferase; U/L, units/liter.
Altogether, our data showed that the relative abundance of the microbiota in Mexican mestizo patients with cirrhosis was significantly different in comparison with the control subjects that had no history of or presented with overt liver disease. In addition, the information enabled a potential bacterial signature directly associated with liver failure to be established.
DiscussionThe most recent studies on microbiota diversity in patients with liver disease indicate that it plays a principal role in the pathophysiology and progression of the pathology.19 However, a large part of those studies have been conducted on populations whose ethnic origins are different from that of the Mexican population. At our laboratory, we believe that is it necessary to obtain local information to find out whether the microbial diversity in Mexican cirrhotic patients is similar to or different from that of other populations. To answer that question, we recruited a pilot group of patients of Mexican mestizo origin that presented with liver disease in the phase of cirrhosis. The majority of the patients recruited were diagnosed with cirrhosis due to C virus infection and were in the compensated phase of the disease, classified as Child-Pugh class A.13 Therefore, complications such as ascites and bleeding of esophageal varices, as well as the ingestion of medications that could modify the composition of the microbiota, were largely controlled and prevented from interfering with the various test results.
Studies that analyze the composition of the microbiota in patients with cirrhosis have shown that Proteobacteria is one of the groups that increases the most.8 There is a significant elevation in the levels of that group of bacteria when the disease is in the decompensated phase or the hospitalization phase, compared with the compensated phase, making it seem that Proteobacteria play a role in the detrimental physiologic conditions of those patients.7 However, in the hospitalization phase, the increase in that taxon could be due to intrahospital infections, which is something that still needs to be determined.
It was interesting to see that, as in other reports, the cirrhotic patients recruited for our study had less bacterial diversity and that they also presented with the same pattern of increased abundance of Proteobacteria (fig. 2), such as Campylobacteraceae, Helicobacteraceae, and Neisseriaceae. In addition, we observed that the order Clostridiales, specifically Veillonella (dispar or parvula), Blautia (obeum or producta), Selenomonas, and Anaerofustis, also had a greater presence in the patients with cirrhosis, compared with the healthy subjects. In addition, that pattern was complemented with data of the correlations between some of the biochemical parameters and the presence of bacteria of the particular genera of Streptococcus, Veillonella, and RF3, all of which were strongly associated with the pathologic state of the patients (fig. 7). In that respect, patients with cirrhosis with advanced grades of damage and whose outcome was liver cancer have been reported to have significant associations between total bilirubin or serum creatinine parameters and the presence of different groups of bacteria, such as Enterobacteria and Clostridium.20
It is also known that different metabolic pathways are enriched in cirrhotic patients, compared with healthy subjects. The pathways of amino acid transport and metabolism, biosynthesis, secondary metabolite transport and metabolism, the production of energy from different carbohydrate sources, and intracellular trafficking and secretion, among others, are enriched, whereas those that decrease include DNA replication, recombination, and repair, to name only a few.4 A more detailed analysis of the metagenome of the microbiota of our patients will establish whether they, too, have alterations in those pathways. It is important to underline that various authors agree that when cirrhotic patients present with problems in protein and carbohydrate metabolism, they have increases in the energy production pathways, in terms of their gut metagenome. It remains to be seen if those pathways are also altered in the population of Mexican mestizo patients with cirrhosis of the liver.
Finally, different articles have shown the usefulness of fecal microbiota transplant in patients affected by Clostridium difficile,21–23 and the results of those interventions appear to have an acceptable degree of success. Nevertheless, there is intense debate as to the possible intermediate and long-term effects, which are still in the testing phases in different clinical trials.24,25 Therefore, other researchers have opted for the use of probiotics (living microorganisms), especially from the genus Lactobacillus. Some of the findings from clinical trials indicate that the administration of Bulgaricus, Bifidobacterium, Plantarum, E. coli Nissle, Bifidobacterium longum, Lactobacillus casei or rhamnosus has positive effects in reducing some of the parameters used for determining liver damage, such as ALT, AST, GGT, and LPS.26 Thus, probiotic and prebiotic consumption has been shown to be a simple and low-cost strategy that, on the one hand, increases the mass of «benign» bacteria that displace the pathogens from the niche that is the gastrointestinal tract, and on the other hand, increases the production of metabolites that contribute to obtaining energy, or more efficient energy, from the foods that affected patients consume.
In the present study, we determined the changes in the relative abundance of intestinal bacteria in a sample of Mexican mestizo cirrhotic patients, compared with a group of subjects with no liver disease. Our data showed that the most abundant groups of intestinal bacteria found in the cirrhotic patients were Proteobacteria and Clostridiales and that they also contributed to associations between some of the biochemical parameters and etiology.
A novel aspect of our study was that we found, with good confidence levels, that some of the taxa identified coincided with those reported by other research groups in regions outside of Mexico. The present analysis is one of the first reports to describe the microbiota in Mexican mestizo patients with cirrhosis of the liver. The results of this study will be complemented a posteriori with the results of the analysis of the oral microbiota of the same group of patients and others that are about to be recruited.
Ethical disclosuresProtection of human and animal subjectsThe authors declare that no experiments were performed on humans or animals for this study.
Confidentiality of data.The authors declare that no patient data appear in this article.
Right to privacy and informed consentThe authors declare that no patient data appear in this article.
Financial disclosureThe present study was partially funded by the Department of Gastroenterology of the INCMNSZ, through the Mexican Public Health Ministry. I. Escalona is the recipient of a grant from the post-graduate scholarship program of excellence of the CONACyT.
Conflict of interestThe authors declare that there is no conflict of interest.
The authors wish to thank A.B. Riveros and C.C. Ruíz for their technical assistance in determining the analytes in COBAS C111, I Medina, MSc, for the discussion in the statistics section, and Dr. Luis F. Uscanga Domínguez for his unconditional support in the realization of this study.
Please cite this article as: Pérez-Monter C, Escalona-Nandez I, Estanes-Hernández A, Noriega-López LG, Torre-Delgadillo A. Determinación de la microbiota intestinal en pacientes cirróticos de población mestizo-mexicana. Revista de Gastroenterología de México. 2019;84:26–35.